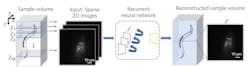
A recurrent neural network is giving a boost to fluorescence samples with a framework for 3D imaging. Researchers at the University of California Los Angeles (UCLA) have created the network, which demonstrates a deep learning-enabled volumetric microscopy framework.
According to the researchers, applications in biomedical and related sciences are increasingly finding benefits in rapid 3D microscopic imaging of fluorescent samples. However, a single 2D image provides a limited axial range, prompting the often-time-consuming process of mechanical scanning of samples using a dense sampling grid for imaging fluorescent samples in 3D. That approach also presents additional light exposure, which “might be toxic and cause unwanted damage such as photobleaching.”
The newly developed technique allows a 3D image to be reconstructed using just a few 2D images of a sample, “providing ~30-fold reduction in the number of scans required to image a fluorescent volume.” The recurrent neural network driving this 3D fluorescence imaging method “intuitively mimics the human brain in processing information and storing memories”—it consolidates important object information and features that frequently appear, while forgetting, or ignoring altogether, some of the redundant information.
In their study, the researchers were able to incorporate spatial features from multiple 2D images to “rapidly reconstruct its 3D fluorescence image.” The volumetric imaging framework was demonstrated using fluorescent C. elegans samples—commonly, those serve as models in biology and bioengineering, the researchers noted.
The UCLA team’s developments in this area are promising in higher imaging speeds for observing 3D specimens “while also mitigating photobleaching- and phototoxicity-related challenges that are frequently observed in 3D fluorescence imaging experiments of live samples.” Reference: L. Huang, H. Chen, Y. Luo, Y. Rivenson, and A. Ozcan, Light Sci. Appl., 10, 62 (2021); https://doi.org/10.1038/s41377-021-00506-9.